Sequential activation of transcriptional programs during development is essential to convert a zygote into a complex organism. A class of cis-regulatory elements called enhancers, which are able to enhance target gene transcription at distances ranging from several to, in rare case, thousands of kilobases, are crucial for precise gene regulation. Enhancers can exist in multiple states, including primed, poised, active, and silenced states. Whereas many excellent reviews have covered how enhancers are activated during cell differentiation and development, enhancer silencing is not yet fully understood.
Prof. Wei Xie's group from the School of Life Sciences, Tsinghua University published a review titled "Activation, decommissioning, and dememorization: enhancers in a life cycle" in Trends in Biochemical Sciences on May 21, 2023. They reviewed the current understanding of enhancer decommissioning and dememorization during development, focusing on their epigenetic states, underlying mechanisms, and potential functions. Finally, they extended these discussions to their involvement in epigenetic inheritance across generations.
Upon silencing of cell type-specific transcriptional programs during development and cell differentiation, active enhancers must be silenced, resulting in enhancer decommissioning. This involves multiple steps including loss of transcription factor (TF) and coactivator binding, histone deacetylation, and in some cases demethylation of H3K4. Despite the removal of characteristic epigenetic marks, decommissioned enhancers do retain some epigenetic signatures, particularly intermediate DNA methylation. These decommissioned enhancers are then fully stripped of epigenetic memories before or after fertilization, which is termed enhancer dememorization. Dememorized enhancers were defined as those that have lost most, if not all, of the epigenetic signatures that are commonly examined. Dememorized enhancers then enter a new life cycle where some are reactivated by embryonic programs, accompanied by resetting of the developmental clock and the start of a new life.
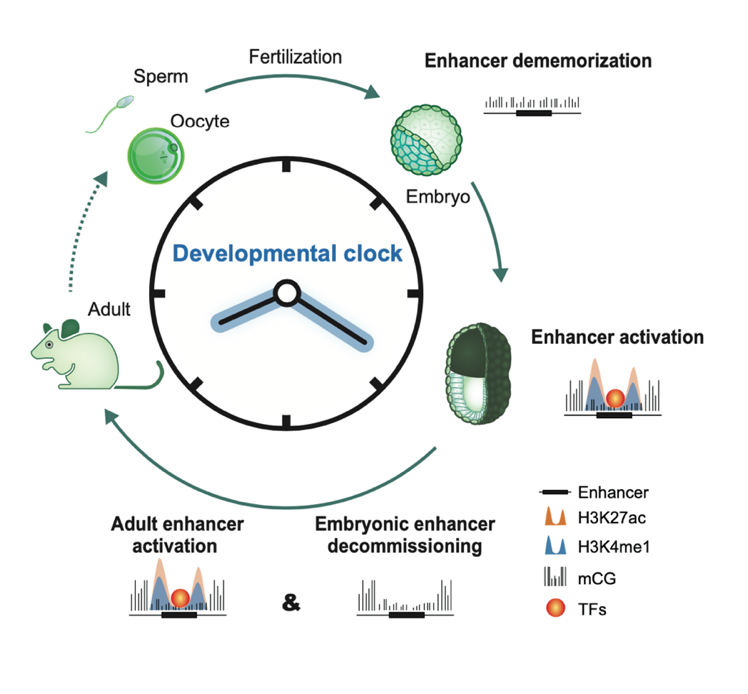
Activation, decommissioning, and dememorization of enhancers in the life cycle
Why decommissioned enhancers are not remethylated is not fully understood because such hypomethylation appears to pose risks of spurious gene activation. One possibility is that decommissioned enhancers could be repurposed in subsequent development to activate the same or different gene targets, and DNA hypomethylation may facilitate their timely reactivation in response to developmental cues. Global epigenetic reprogramming, which is essential to convert gametes into totipotent embryos, removes parental memories carried by epigenetic marks and establishes embryo-specific epigenomes. One prominent feature of enhancer dememorization is that it often occurs almost promiscuously at nearly all enhancers, and such effects are executed presumably by altering the expression or global actions of the corresponding epigenetic factors. By contrast, enhancer decommissioning typically entails the actions of epigenetic factors at individual loci.
In summary, epigenetic signatures of regulatory elements not only reflect past memory of transcription but also retain the potential for future activation. Deciphering how enhancers are silenced is equally important for understanding how cell fate is dynamically determined and reset. Enhancer decommissioning effectively terminates transcription upon changes in developmental cues to allow activation of the transcriptional programs for the next stage. Enhancer dememorization then ensures the effective erasure of parental memories, thus achieving the resetting of the epigenetic clock.
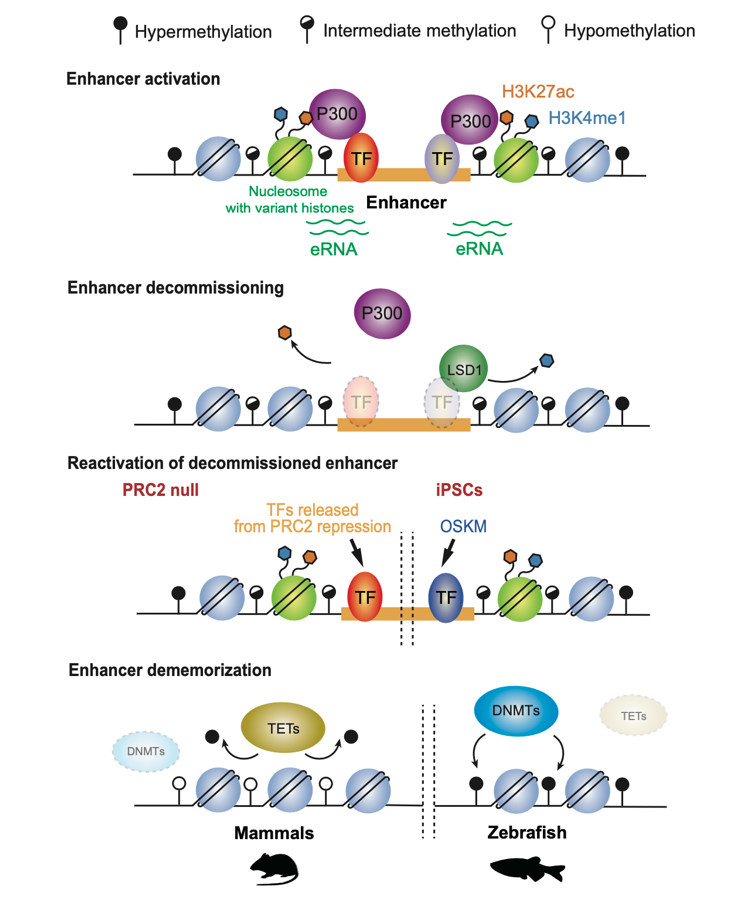
Regulatory mechanisms underlying enhancer decommissioning and dememorization
Prof. Wei Xie from the School of Life Sciences, Tsinghua University is the corresponding author of this paper. Research Assistant Professor Xiaotong Wu from the School of Life Sciences, Tsinghua University is the first author of this paper. Ph.D. student Xi Wu also provided important help for the writing of the paper. The project was supported by the National Key R&D Program of China, the National Natural Science Foundation of China, and the Tsinghua-Peking Center for Life Sciences. Wei Xie is a New Cornerstone Investigator and a Howard Hughes Medical Institute (HHMI) International Research Scholar.
Original article link: https://doi.org/10.1016/j.tibs.2023.04.005
Editor: Li Han

